
In order to measure volumes or to repeat measurements across the entire stack, some simple macro programming is required. ImageJ has numerous tools for measuring objects from images, however most of these tools are built to process a single image and return a single result. Measurement of Object Volume from Image Stacks Select the image acquisition you would like to load, and click OK. Please note that Bioformats will load each checked image set into memory, and if image stacks are each very large, it may consume a lot of memory on your system, which can seriously effect the performance of your computer. If the proprietary image format stores multiple image sets in the same file, the Bioformats plugin will show you an additional dialog allowing you to select which image acquisitions you would like to load. Select all required settings and then click OK. This can be accomplished by selecting the “Hyperstack” option from the “View stack with” dropdown box at the top left of the dialog. For multichannel images, it is convenient to load the stack as a hyperstack, which will automatically separate channel frames and provide a slider bar to select the channel of interest. The first dialog screen (Figure 3) allows you to modify various settings about how Bioformats will load the images from the proprietary format, for example, you can tell Bioformats to split the channels into individual stacks (Split channels checkbox), or to automatically stitch tiles of a multi-field image together (Stitch tiles checkbox). Bioformats is a powerful plugin released by the University of Dundee and Open Microscopy Environment, and enables loading of dozens of proprietary image formats directly into ImageJ. czi files) can be imported into ImageJ by using the File > Import > Bioformats command (Fiji has Bioformats installed by default). Proprietary image formats (such as the Leica. 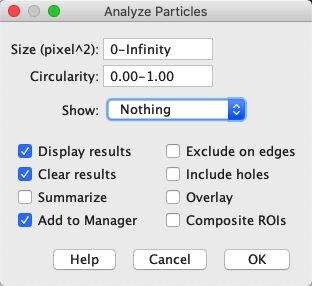
TIF hyperstack, proprietary formats such as Leica. TIF files) where each image is a single z-plane in a single color or as a single file containing all images in the stack (e.g. Depending on the type of instrument and settings used during acquisition, the image may be a sequence of individual images (typically as. Three-dimensional confocal imaging results in a collection of images as planes incrementing along the Z axis, known as an “image stack” (also known as a “z-stack”). Loading and Processing Image Stacks in ImageJ (or Fiji) In conjunction with fluorescent labeling techniques, tissue clearing dramatically increases the depths at which confocal microscopy can image into tissue, allowing for large scale reconstructions of tissue histology and morphology on large and thick pieces of tissue (> 1 mm). Visikol HISTO, CLARITY, 3DISCO) can by employed to render tissue specimens optically transparent. To increase the achievable depth at which confocal microscopes can be utilized to obtain images within tissues, clearing techniques (e.g. For deep tissue imaging (> 2mm), the use of glycerol or water immersion objectives is preferred. For most applications, air objectives work effectively the highest numerical aperture (NA) available at the magnification required should be selected, as higher NA results in more light entering the objective, which yields higher contrast and signal. The selection of microscope objective for imaging can have serious impacts on quality of images obtained. This is commonly known as “optical sectioning” wherein thick tissue sections are employed and utilizing the optical ability of the confocal microscope as an optical “microtome,” structural details of the tissue can be revealed in three-dimensions. In a fixed position, providing a three-dimensional sampling of the tissue being examined.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
